Data association implementation that uses the Malhanobis Distance only as criterion to make observation matches.
More...
#include <jcbb.hpp>
|
| void | search_branch_and_bound (int current_obs_idx, int num_observations, int num_landmarks, const Eigen::MatrixXd &distances, Eigen::VectorXi ¤t_hypothesis, Eigen::VectorXi &best_hypothesis, double &best_score, double current_score) const |
| | Recursive branch and bound search to find the best hypothesis.
|
| |
| bool | is_landmark_already_assigned (const Eigen::VectorXi &hypothesis, int landmark_idx) const |
| | Check if a landmark is already assigned in the current hypothesis.
|
| |
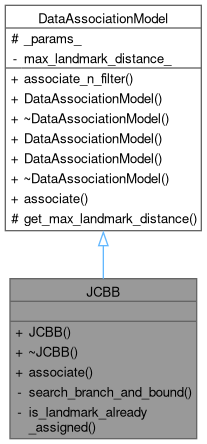
Data association implementation that uses the Malhanobis Distance only as criterion to make observation matches.
Definition at line 17 of file jcbb.hpp.
◆ JCBB()
◆ ~JCBB()
◆ associate()
| Eigen::VectorXi JCBB::associate |
( |
const Eigen::VectorXd & |
landmarks, |
|
|
const Eigen::VectorXd & |
observations, |
|
|
const Eigen::MatrixXd & |
covariance, |
|
|
const Eigen::VectorXd & |
observation_confidences |
|
) |
| const |
|
overridevirtual |
Perform data association using the Joint Compatibility Branch & Bound algorithm.
This function finds the most consistent assignment of 2D observations to known 2D landmarks by maximizing the number of mutually compatible pairs. Uses Euclidean distance as the compatibility measure and a branch & bound search to efficiently explore the space of hypotheses.
- Parameters
-
| landmark_positions | Landmark positions in format [x1, y1, x2, y2, ...]. |
| observation_positions | Observation positions in format [x1, y1, x2, y2, ...]. |
| covariance | Covariance matrix of state estimate (unused in this version). |
| observation_confidences | Confidences for each observation (unused in this version). |
- Returns
- Eigen::VectorXi of size num_observations:
- If element i >= 0 → index of associated landmark.
- If element i == -1 → observation is unassigned (new/clutter).
Implements DataAssociationModel.
Definition at line 3 of file jcbb.cpp.
◆ is_landmark_already_assigned()
| bool JCBB::is_landmark_already_assigned |
( |
const Eigen::VectorXi & |
hypothesis, |
|
|
int |
landmark_idx |
|
) |
| const |
|
private |
Check if a landmark is already assigned in the current hypothesis.
- Parameters
-
| hypothesis | Current hypothesis vector. |
| landmark_idx | Index of the landmark to check. |
- Returns
- true if landmark is already assigned, false otherwise.
Definition at line 84 of file jcbb.cpp.
◆ search_branch_and_bound()
| void JCBB::search_branch_and_bound |
( |
int |
current_obs_idx, |
|
|
int |
num_observations, |
|
|
int |
num_landmarks, |
|
|
const Eigen::MatrixXd & |
distances, |
|
|
Eigen::VectorXi & |
current_hypothesis, |
|
|
Eigen::VectorXi & |
best_hypothesis, |
|
|
double & |
best_score, |
|
|
double |
current_score |
|
) |
| const |
|
private |
Recursive branch and bound search to find the best hypothesis.
- Parameters
-
| current_obs_idx | Index of the observation being processed. |
| num_observations | Total number of observations. |
| num_landmarks | Total number of landmarks. |
| distances | Precomputed Euclidean distances between observations and landmarks. |
| current_hypothesis | Current hypothesis (partial). |
| best_hypothesis | Best hypothesis found so far (output). |
| best_score | Best score found so far (output). |
| current_score | Score of the current partial hypothesis. |
Definition at line 33 of file jcbb.cpp.
The documentation for this class was generated from the following files:
- src/perception_sensor_lib/include/perception_sensor_lib/data_association/jcbb.hpp
- src/perception_sensor_lib/src/data_association/jcbb.cpp
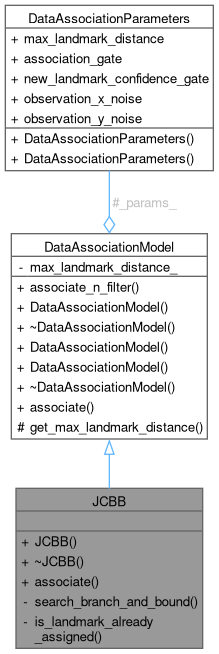


 Public Member Functions inherited from DataAssociationModel
Public Member Functions inherited from DataAssociationModel Protected Member Functions inherited from DataAssociationModel
Protected Member Functions inherited from DataAssociationModel Protected Attributes inherited from DataAssociationModel
Protected Attributes inherited from DataAssociationModel