Loading...
Searching...
No Matches
MaximumLikelihoodMD Class Reference
Data association implementation that uses the Malhanobis Distance only as criterion to make observation matches. More...
#include <maximum_likelihood_md.hpp>
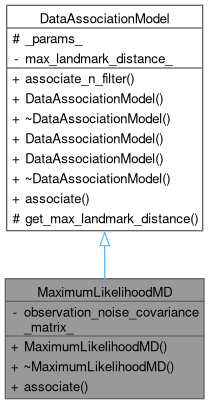
Inheritance diagram for MaximumLikelihoodMD:

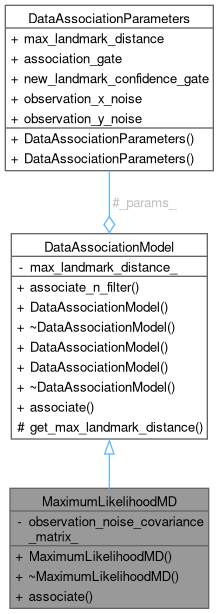
Collaboration diagram for MaximumLikelihoodMD:

Public Member Functions | |
| MaximumLikelihoodMD (const DataAssociationParameters ¶ms) | |
| ~MaximumLikelihoodMD ()=default | |
| Eigen::VectorXi | associate (const Eigen::VectorXd &landmarks, const Eigen::VectorXd &observations, const Eigen::MatrixXd &covariance, const Eigen::VectorXd &observation_confidences) const override |
| This function associates the landmarks with the observations. | |
 Public Member Functions inherited from DataAssociationModel Public Member Functions inherited from DataAssociationModel | |
| virtual int | associate_n_filter (const std::vector< common_lib::structures::Cone > &perception_map, Eigen::VectorXf &_x_vector_, Eigen::MatrixXf &_p_matrix_, std::vector< int > &matched_ids, std::vector< Eigen::Vector2f > &matched_cone_positions, std::vector< Eigen::Vector2f > &new_features, ObservationModel *observation_model) const =0 |
| Associate the observed landmarks to the expected landmarks and update the state vector and the covariance matrix. | |
| DataAssociationModel (float max_landmark_distance) | |
| virtual | ~DataAssociationModel ()=default |
| DataAssociationModel ()=default | |
| DataAssociationModel (DataAssociationParameters params) | |
| virtual | ~DataAssociationModel ()=default |
Private Attributes | |
| Eigen::Matrix2d | observation_noise_covariance_matrix_ |
Additional Inherited Members | |
 Protected Member Functions inherited from DataAssociationModel Protected Member Functions inherited from DataAssociationModel | |
| float | get_max_landmark_distance () const |
 Protected Attributes inherited from DataAssociationModel Protected Attributes inherited from DataAssociationModel | |
| DataAssociationParameters | _params_ |
Detailed Description
Data association implementation that uses the Malhanobis Distance only as criterion to make observation matches.
Definition at line 13 of file maximum_likelihood_md.hpp.
Constructor & Destructor Documentation
◆ MaximumLikelihoodMD()
|
inline |
Definition at line 17 of file maximum_likelihood_md.hpp.
◆ ~MaximumLikelihoodMD()
|
default |
Member Function Documentation
◆ associate()
|
overridevirtual |
This function associates the landmarks with the observations.
- Parameters
-
landmarks Landmarks in the form of [x1, y1, x2, y2, ...] in the global frame observations Observations in the form of [x1, y1, x2, y2, ...] in the global frame covariance Covariance matrix of the landmark vector observation_confidences Confidence in the observations in the same order as the observations
- Returns
- Eigen::VectorXi Each entry corresponds to an observation and contains the index of the landmark that the observation is associated with in the landmark vector (x coordinate). If the observation is considered new, the entry is -1. If the observation is considered an outlier, the entry is -2.
Implements DataAssociationModel.
Definition at line 9 of file maximum_likelihood_md.cpp.
Here is the caller graph for this function:
Member Data Documentation
◆ observation_noise_covariance_matrix_
|
private |
Definition at line 14 of file maximum_likelihood_md.hpp.
The documentation for this class was generated from the following files:
- src/perception_sensor_lib/include/perception_sensor_lib/data_association/maximum_likelihood_md.hpp
- src/perception_sensor_lib/src/data_association/maximum_likelihood_md.cpp